Literature Review
Epilepsy affects over 50 million people worldwide, and about 30% of patients suffer from drug-resistant epilepsy, for whom surgery is often the only curative option. The success of surgical intervention depends critically on the accurate identification of the epileptogenic zone (EZ), specifically the Seizure Onset Zone (SOZ), the region where seizures originate.
Stereo-electroencephalography (sEEG), which involves intracranial recordings from depth electrodes, has become a gold standard for delineating the SOZ due to its ability to capture high-resolution spatiotemporal activity.
Key Biomarkers for SOZ Detection
- High Frequency Oscillations (HFOs): 80-500 Hz signals considered reliable markers
- Interictal Spikes: Interictal epileptiform discharges remain standard clinical markers
- Phase-Amplitude Coupling (PAC): Captures cross-frequency interactions
- Connectivity: Functional connectivity adds valuable information for SOZ localization
Traditional approaches rely heavily on expert visual review, which is time-consuming, subjective, and prone to inter-rater variability. Machine learning methods including CNNs, RNNs, GNNs, and Transformers have shown promise in automating this process.
Dataset Description
We will employ four multi-center datasets to ensure variability in acquisition protocols, electrode configurations, and patient demographics. This heterogeneity will allow us to test the generalizability of our models across centers and clinical settings.
De-identified patient data (n=58) containing electrophysiologic data for interictal and ictal periods from the Hospital of the University of Pennsylvania.
View DatasetDe-identified patient data (n=39) containing iEEG recordings with sleep/wake state annotations from multiple centers including NIH, Johns Hopkins, and University of Miami.
View DatasetDe-identified patient data (n=100) containing iEEG and EEG data from 5 centers, with 4 centers' data publicly available.
View DatasetIntracranial EEG recordings from patients undergoing pre-surgical evaluation. This dataset is not publicly available but will be used for training.
Problem Definition and Motivation
For patients with drug-resistant focal epilepsy, surgical resection of the SOZ can be curative. However, the main clinical challenge lies in accurately identifying the SOZ from sEEG recordings. Mislocalization leads to failed surgeries and persistent seizures.
Current manual approaches are inefficient and limited by subjective interpretation. Our motivation is to develop machine learning models that can automatically detect the SOZ with high accuracy, robustness across datasets, and interpretability aligned with known biomarkers.
The project's impact extends beyond accuracy: automating SOZ detection can reduce diagnostic time, improve consistency, and potentially expand surgical candidacy by making evaluation more scalable.
Proposed Solutions
Preprocessing Pipeline
We adopt a multi-pronged preprocessing pipeline combining time-frequency decomposition with clinically validated biomarkers:
Five levels of decomposition with Daubechies-4 basis to generate localized time-frequency features for each channel.
Automated HFO detection (80-500 Hz) from WPT representation provides features sensitive to microstructural pathological activity.
Interictal epileptiform discharges are standard clinical markers of epileptogenic zones.
Functional connectivity measures including Amplitude Envelope Correlation (AEC) and Phase-Locking Value (PLV).
Machine Learning Models
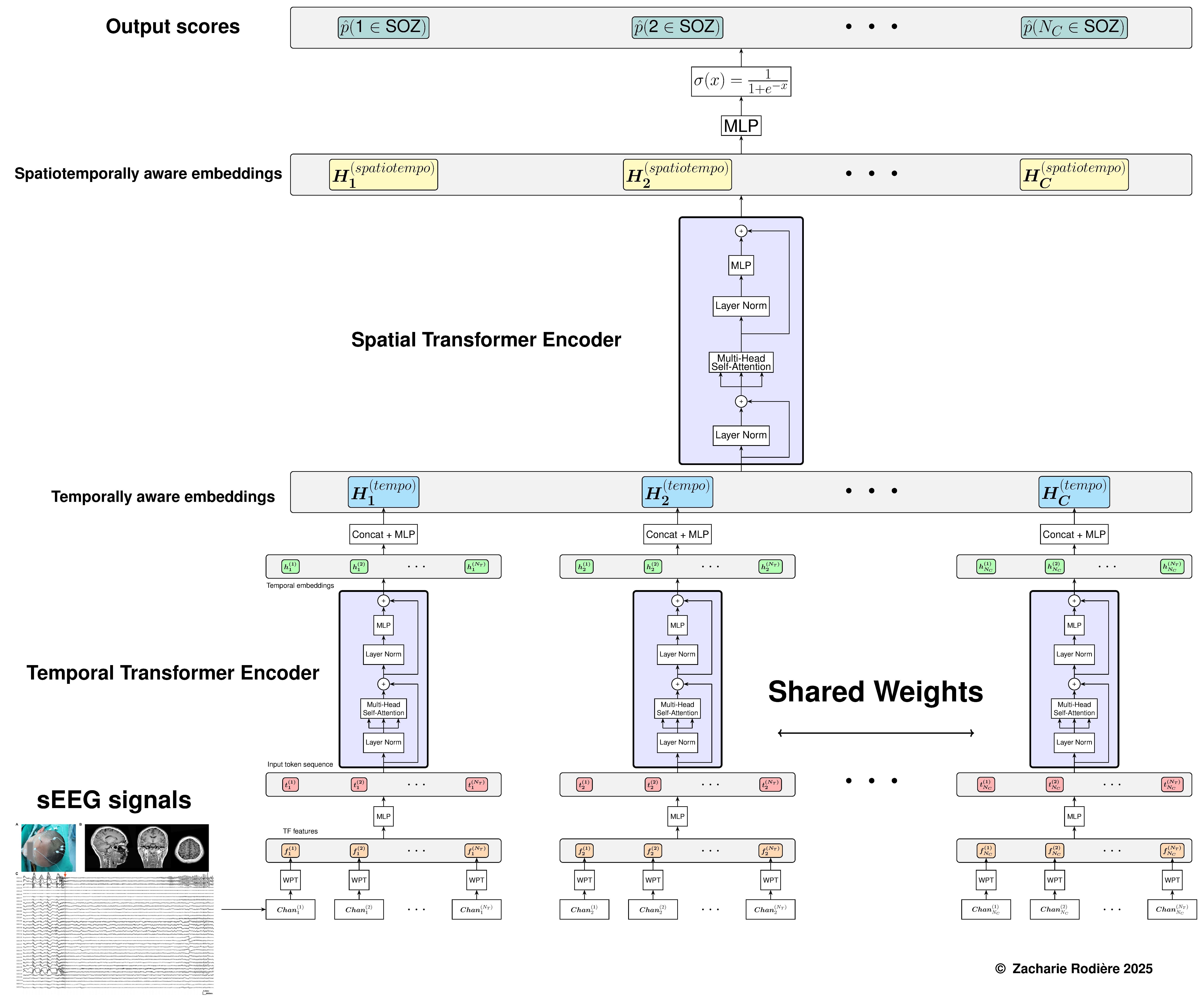
Learns channel-wise temporal representations with spatial attention module to highlight inter-channel relationships. Uses pretraining via spatial contrastive learning.
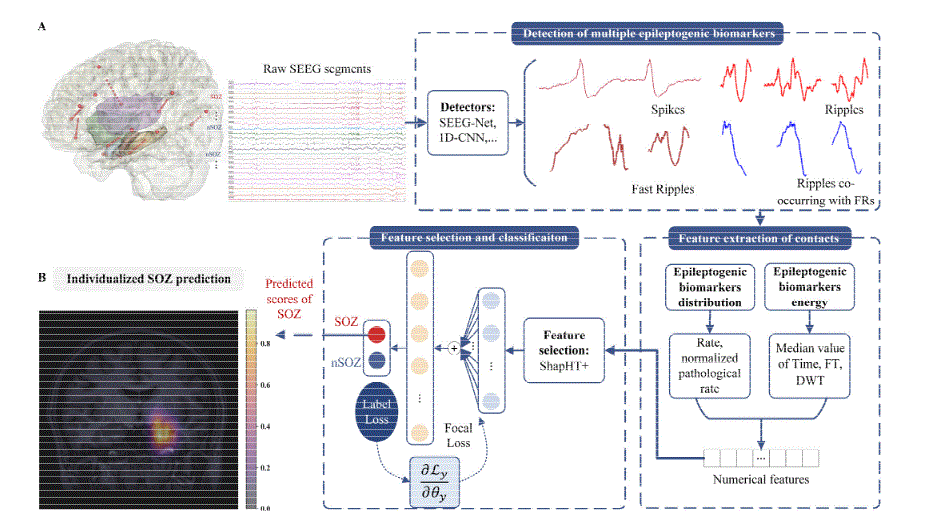
Reproduction of a recent deep learning approach that utilizes multiple feature types and claims to achieve strong performance on seizure onset zone detection. We aim to reproduce and validate this methodology on our multi-center datasets.
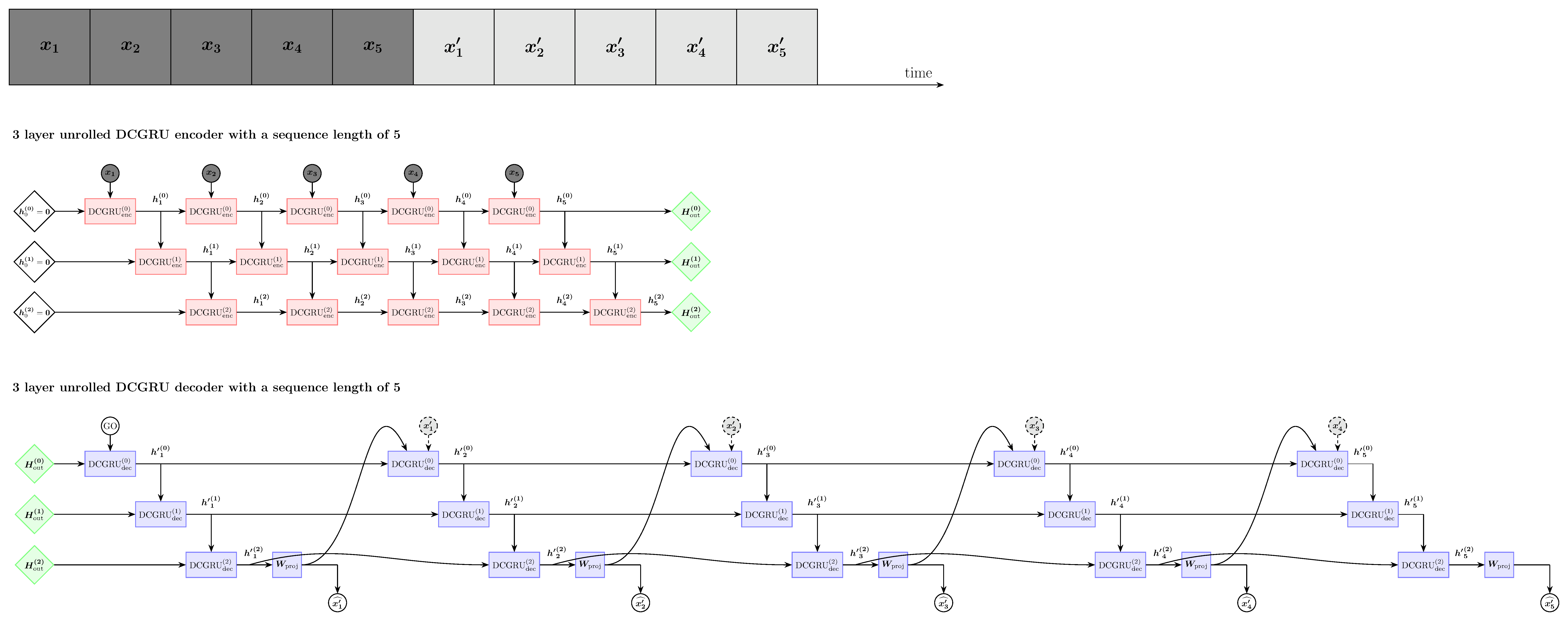
Self-supervised exploration using DCGRU to model the graph structure of sEEG electrodes. This approach learns signal representations that may enable anomaly detection on learned features if we develop a robust model of normal signal patterns.
Expected Results and Evaluation
Evaluation Metrics
Since the SOZ is a physical region in the brain but we can only assign binary labels to each electrical contact, evaluation will be performed in terms of channel-wise binary classification:
Expected Performance
- Transformer model: Highest AUROC (≥0.80)
- Multi-Feature Model: Reproduction of published results with validation on our datasets
- DCGRU (Self-supervised): Exploratory approach for anomaly detection on learned features
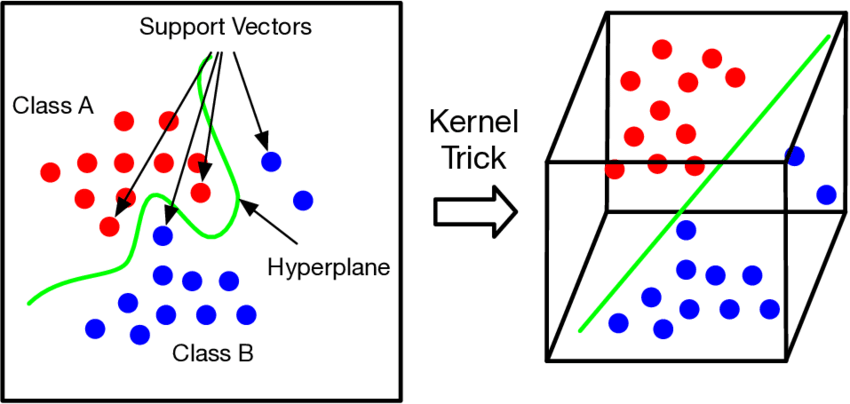
- SVM: Baseline performance (AUROC ~0.60-0.65)
By triangulating results across models and datasets, we aim to validate the robustness of our approach and ensure consistent performance with minimal per-patient variability.
Team Contributions
Project Timeline - Gantt Chart
Our project timeline showing the planned phases, milestones, and team responsibilities throughout the semester.
Project Award Eligibility
🏆 Outstanding Project Award Consideration
We would like to opt-in to be considered for the "Outstanding Project" award. Our team is committed to delivering high-quality research that advances the field of automated seizure onset zone detection using machine learning.
We believe our comprehensive approach combining multiple datasets, advanced preprocessing techniques, and diverse machine learning architectures positions us well for this recognition. Our project addresses a critical clinical need and demonstrates both technical innovation and practical impact.